This post accompanies the CHIMIA Chemical Education Highlight DIY a molecule: Generative models in chemistry. To jump straight to hands-on molecule generation, open the GenMol Colab notebook.
TL;DR
- Generative models propose new molecules with desired structures and properties. It is even possible to incorporate experiments as feedback and properties as “conditioning signals” that steer a model.
- GenMol treats molecule design as a “fill-in-the-blank problem” over molecular fragments.
- You can try it yourself, right now, in the browser with no installation needed. If you prefer to install the code yourself, access the code repository here.
What is a generative model, and why should chemists care?
Most chemists have encountered predictive models: you feed in a structure, and the model returns a property such as logP, binding affinity. These are useful, but they answer a fixed question: given this molecule, what is its property?
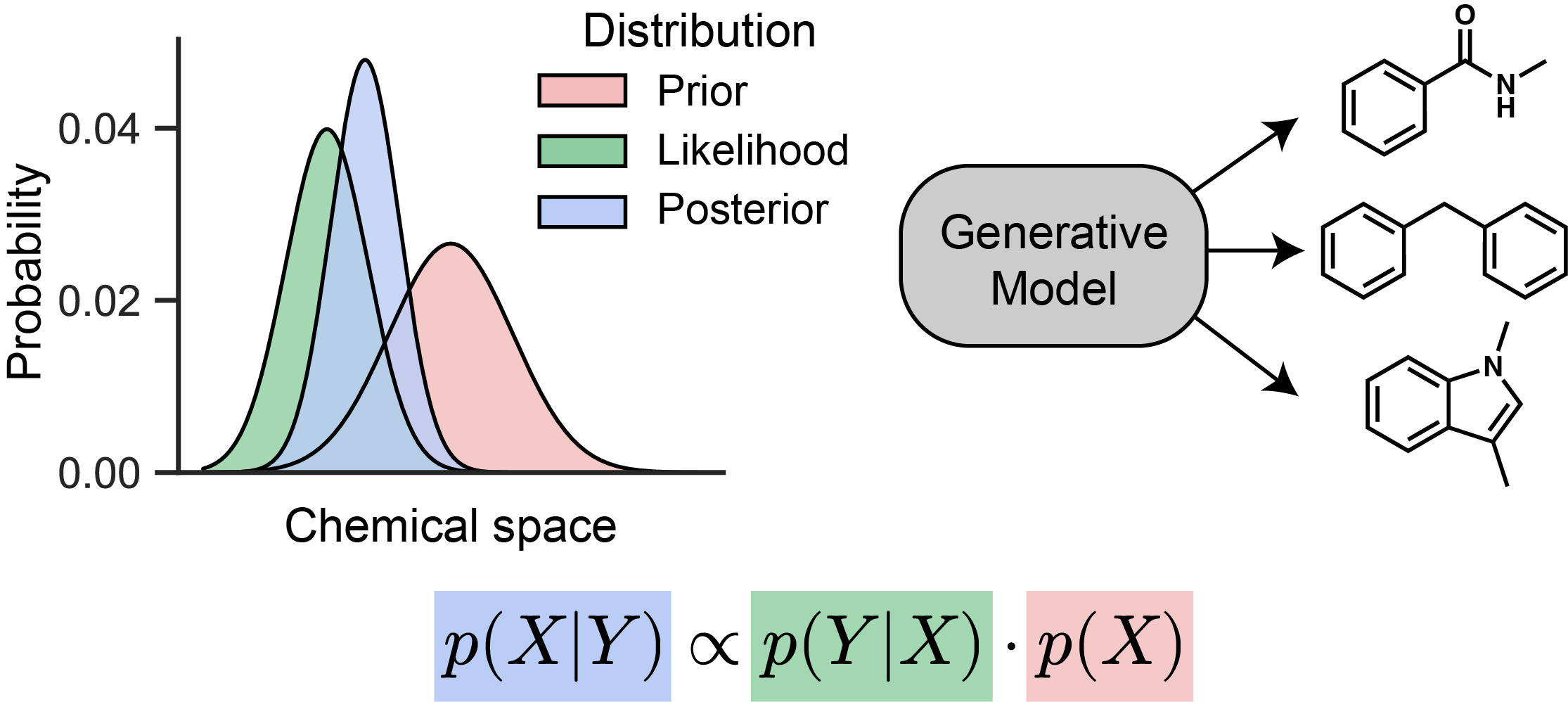
A generative model asks something different. Rather than mapping structures to labels, it learns the distribution of molecules themselves — what drug-like compounds tend to look like, for example — and draws new samples from it. The result is a model that can propose structures.
The more useful version of this is conditional generation: given a desired property or structural constraint, generate molecules that satisfy it. This is where generative models connect directly to how medicinal chemists work.
Molecules as fragments, generation as fill-in-the-blank
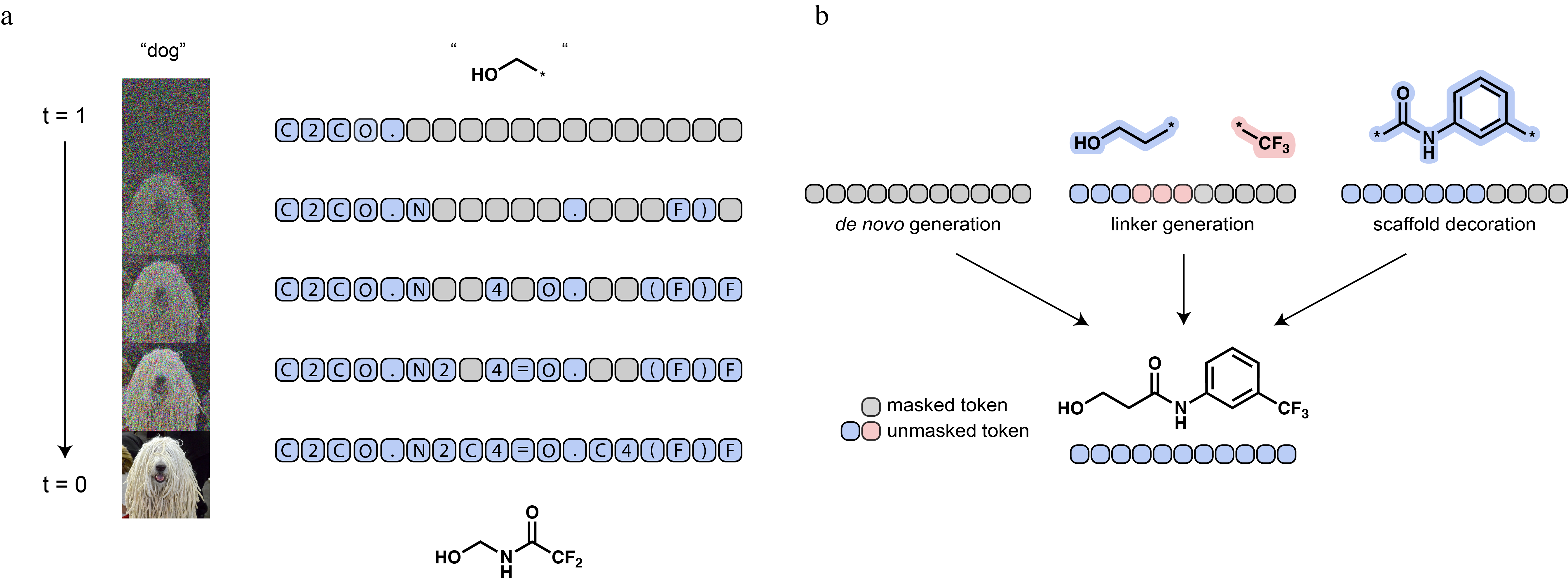
GenMol [1] uses the SAFE molecular representation [2], which breaks a molecule into its constituent fragments (such as ring systems, chains, functional groups) and treats them as the basic units of generation.
With this, generating a molecule becomes a fill-in-the-blank problem over fragments. The mechanism is called masked diffusion: during training, fragments are progressively replaced with a [MASK] token until the whole molecule is hidden. The model learns to reverse this, recovering the structure. At generation time, it starts from a fully masked sequence and iteratively fills in fragments until a complete molecule emerges.
What makes this particularly suited to drug discovery is that any part of the molecule can be fixed as context while the rest is generated. Three tasks, that we will explore in the accompanying notebook, fall out naturally from the same model:
| Task | Fixed (context) | Generated |
|---|---|---|
| De novo generation | nothing | entire molecule |
| Scaffold decoration | core ring system | side chains |
| Linker design | two terminal fragments | the bridge between them |
Try it yourself
The companion Colab notebook runs GenMol in the browser on a free GPU.
You can explore the following:
- Generating drug-like molecules from scratch
- Decorating a scaffold you draw directly in the notebook
- Connecting two fragments with a generated linker
- Seeing where your molecules land in chemical space, projected onto a PubChem reference map
Three parameters are worth exploring. Temperature controls how closely the model stays to its training distribution — low values produce conventional, drug-like structures; high values push into less typical territory. Randomness controls the order in which fragments are unmasked, adding a second source of variation. Gamma, which is available for scaffold decoration and linker design, controls how strongly the model is guided by the provided fragment: higher values push the output to more faithfully incorporate the given scaffold or fragment, at the cost of some diversity. Zero disables it entirely. I highly encourage you to explore the effects of tuning those parameters yourself!
Note that generative models can propose structures that are difficult or impossible to make, and many outputs will be close (or exact) variants of known compounds rather than genuinely novel scaffolds. These tools are most useful when used with chemical judgment — so test the limits, pay attention to where the model fails and treat its proposals as starting points rather than answers. Happy creating!
References
[1] M. Lederbauer, CHIMIA, 2026. doi: 10.2533/chimia.2026.335
[2] S. Lee, K. Kreis, S. Prasad Veccham, M. Liu, D. Reidenbach, Y. Peng, S. Paliwal, W. Nie, A. Vahdat, ICML, 2025. doi: 10.48550/arXiv.2501.06158
[3] Noutahi, C. Gabellini, M. Craig, J. S. C. Lim, P. Tossou, Dig. Disc., 2024, 3, 796–804. doi: 10.1039/D4DD00019F
Citation
@online{lederbauer2026,
author = {Lederbauer, Magdalena},
title = {DIY a Molecule: {Generative} Models in Chemistry},
date = {2026-03-31},
url = {https://mlederbauer.com/posts/2026-03-31-genai-fbdd/},
langid = {en}
}